When I ship a medical AI system into a regulated environment, the single biggest predictor of downstream pain isn’t the model architecture, it’s the dataset. Whether I’m validating hallucination rates in an LLM-based CDS tool or stress-testing a sepsis model against edge cases, the choice of medical AI datasets determines what I can credibly claim to regulators, clinicians, and my own QA team.
In this guide, I’ll walk through the major open clinical, imaging, and omics datasets I actually use or recommend, from MIMIC and eICU to CheXpert, GTEx, and more, along with practical notes on access, compliance, and limitations. I’ll focus on how these datasets map to real deployment questions under HIPAA/GDPR rather than abstract benchmark bragging rights.
Medical disclaimer: Nothing here is medical advice or a substitute for clinical judgment. Use these datasets for research and development only, and always follow your local IRB, institutional, and regulatory requirements.
Table of Contents
Comprehensive Overview of MIMIC-III and Clinical Text Datasets in Medical AI
In-Depth Overview of MIMIC-III ICU Records for Medical Research and AI Development
MIMIC-III (v1.4) from PhysioNet is still my default recommendation when someone asks, “Where do I start with medical AI datasets for EHR data?” It contains de-identified ICU data for over 40,000 patients at Beth Israel Deaconess (Johnson et al., Sci Data 2016: PhysioNet, Nature article).
From a practical engineering standpoint, I use MIMIC-III for:
- Time-series modeling: vitals, labs, meds for sequence models (e.g., sepsis prediction, AKI, mortality).
- Text modeling: discharge summaries and radiology reports for note classification, entity extraction, and LLM evaluation.
- Reproducible baselines: it’s the dataset underlying dozens of benchmark papers, so it’s easy to compare against published work.
In one ICU decision-support prototype I worked on, I first tuned a risk model on MIMIC-III to lock in feature engineering and evaluation protocols (AUROC, calibration, subgroup analyses), then moved to our private dataset only after the pipeline was stable. That separation helped during regulatory review because we could show method development on public data and validation on in-house, PHI-containing data.
Regulatory-wise:
- Data are de-identified under HIPAA Safe Harbor, but you still need to complete the required CITI training and data use agreement (DUA).
- MIMIC-III is single-center, US-based, heavily ICU-biased. I never treat it as a production representativeness proxy, only a development and benchmarking platform.
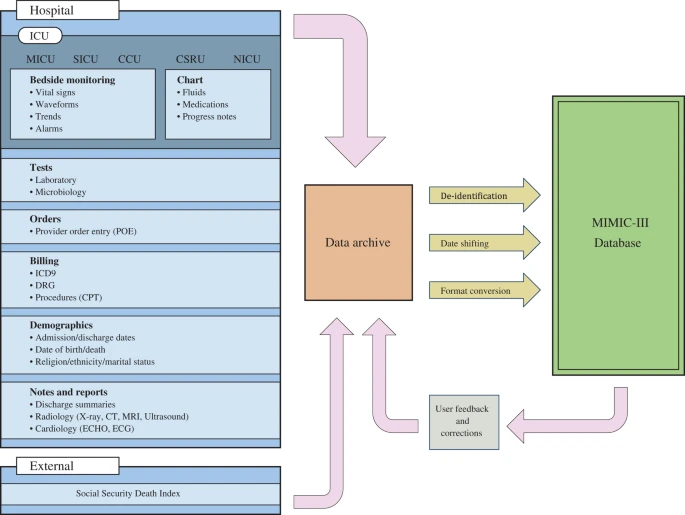
Exploring Other Key Electronic Health Record (EHR) Text Datasets: MIMIC-IV and eICU
For newer work, I increasingly prefer MIMIC-IV (PhysioNet, Johnson et al., Sci Data 2023). It:
- Splits hospital and ICU data more cleanly
- Has updated coding (ICD-9/10), medication, and note structures
- Maintains the same robust de-identification and DUA process
If I’m evaluating LLMs on chart summarization or structured label extraction, MIMIC-IV’s note corpus is simply more aligned with contemporary documentation.
The eICU Collaborative Research Database (eICU-CRD, Pollard et al., Sci Data 2018) fills an important gap:
- Multi-center ICU data from over 200 US hospitals
- Rich time-series plus some free text
In one project examining model robustness, I deliberately trained on MIMIC-III and tested on eICU to approximate domain shift. The drop in calibration across smaller community hospitals was a useful early warning before we ever touched proprietary data.

Key caveats across these EHR datasets:
- They’re ICU-heavy: not great for chronic outpatient trajectories.
- Demographics skew US, with underrepresentation of certain racial and socioeconomic groups.
- All LLM-style work must respect the DUA and avoid any attempt at re-identification.
For HIPAA/GDPR-compliant deployments, I use these only for method development, pre-training, and benchmarking. Final validation must be performed on local, governed datasets with appropriate DPO/IRB oversight.
Key Medical Imaging Datasets for AI Innovation
Accessing Public Chest X-ray and CT Imaging Databases: ChestX-ray14 and CheXpert
For imaging-heavy pipelines, especially when I’m validating triage or QA systems, chest radiograph datasets are the workhorses.
- NIH ChestX-ray14 (NIH CC, Wang et al., 2017) offers >100k frontal chest X-rays with 14 disease labels mined from reports.
- CheXpert (Stanford ML Group, Irvin et al., 2019) contributes >220k chest X-rays with uncertainty-aware labels and a well-defined validation set.
In practice, I use these datasets to:
- Pre-train encoders (CNNs, vision transformers) for transfer learning
- Run benchmark-style evaluations for classification, localization, and label uncertainty
- Stress-test LLM+vision systems that read radiology reports and images jointly
Both datasets rely on NLP-derived labels from reports, which means:
- They’re great for weakly supervised learning.
- They’re not perfect for safety-critical thresholds. For example, mislabelled cardiomegaly or subtle pneumothorax can skew calibration.
Whenever I’ve considered using models trained on these data near clinical workflows, I’ve enforced an additional local fine-tuning and silent shadow deployment phase with clinician review before even thinking about active use.
Exploring Pathology and Ophthalmology Imaging Datasets for AI-driven Diagnosis
For histopathology, the CAMELYON16/17 challenges (Camelyon17, Bándi et al., 2019) remain foundational:

- Whole-slide images for lymph node metastasis detection in breast cancer
- Strongly annotated metastasis regions
I’ve used CAMELYON slides to validate WSIs preprocessing (tiling, color normalization) and to benchmark segmentation/localization models. But I’m careful not to overfit model design to these narrow tasks, they’re highly specific to breast cancer lymph nodes.
For other modalities (retina, CT segmentation, etc.), I rely heavily on datasets hosted via Grand Challenge (grand-challenge.org). These give:
- Public training sets with task-specific annotations
- Hidden test sets and standardized metrics
- Shared leaderboards that are surprisingly useful during internal model selection
Again, these are invaluable for technical benchmarking, not for skipping local validation. Scanner differences, staining protocols, and population variations can be dramatic once you leave the competition sandbox.
Biomedical Literature and Genomic Data for AI Applications
Leveraging PubMed and Other Biomedical Text Corpora for Medical AI Training
When I’m evaluating LLMs for clinical QA, I care a lot about what underlying biomedical text they’ve actually seen.
- PubMed and PubMed Central full-text (via appropriate licenses) remain the backbone for biomedical language understanding.
- For hallucination analysis, I often construct test sets where the ground truth is anchored in peer-reviewed articles or guidelines, then probe whether the model cites or contradicts that evidence.
A practical pattern I like:
- Pre-train or adapt on large biomedical corpora (PubMed, clinical guidelines, FDA labels).
- Fine-tune on de-identified clinical notes (e.g., MIMIC-IV) where permitted.
- Evaluate against a curated benchmark that mixes literature-grounded questions and note-grounded tasks.
This lets me quantify:
- Reference accuracy vs. hallucinations
- Guideline adherence (e.g., NIH, WHO recommendations)
- Ability to state uncertainty instead of fabricating specifics
Key Genomic and Proteomic Datasets Supporting Advances in Medical AI
On the omics side, I’ve found three resources particularly impactful:
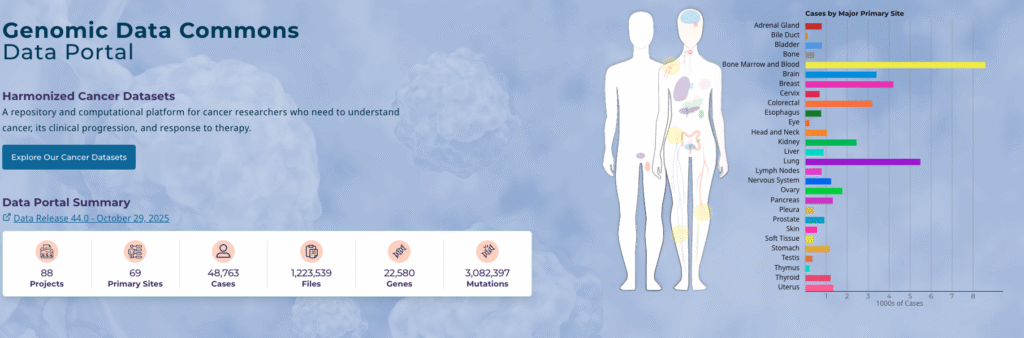
- Genomic Data Commons (GDC) for cancer genomics (portal.gdc.cancer.gov) – integrates TCGA and other programs.
- GTEx for tissue-specific gene expression (gtexportal.org: GTEx Consortium, Nature Genetics 2013+).
- AlphaFold DB for protein structures (alphafold.ebi.ac.uk).
These datasets are ideal for models that:
- Map from variants to predicted phenotypes
- Learn multimodal relationships between imaging, pathology, and genomics
- Support drug discovery workflows
But, I treat these firmly as research tools. Germline and somatic genomic data are highly sensitive: even when access is restricted and controlled, GDPR and local genomics regulations (e.g., in the EU) can be stricter than typical health data rules. Any pipeline touching patient-linked omics always goes through formal governance and explicit consent review.
The Impact of Open Data on Accelerating AI Progress in Healthcare
How Public Medical Datasets Enable Breakthroughs in AI Research and Applications
From my vantage point, open medical AI datasets have changed three things:
- Reproducibility: When a paper reports a sepsis model on MIMIC-IV or a pneumonia detector on CheXpert, I can reproduce it, extend it, and meaningfully compare.
- Talent pipeline: Trainees and engineers outside major academic centers can build serious models without privileged data access.
- Safety culture: It’s easier to experiment with hallucination mitigation, calibration, and robustness on de-identified data before touching PHI.
In one hospital deployment, we used MIMIC/eICU for method development and then ported only the most promising architectures into our governed training environment. That separation substantially reduced review friction with compliance and privacy teams.
Community Challenges and Benchmarks: Collaborative Efforts to Drive Medical AI Innovation
Community challenges, many hosted on Grand Challenge, provide:
- Standardized tasks, metrics, and hidden test sets
- Strong baselines and open-source code
- Clear leaderboards that help cut through marketing claims
Examples range from CAMELYON for pathology to various organ segmentation and detection tasks.
I often re-use challenge leaderboards as sanity checks: if my model underperforms the median of a three-year-old competition, it’s usually not ready for a production validation round.
Still, open benchmarks can become stale. Disease prevalence, scanners, coding schemes, and clinical practice patterns evolve. When using any benchmark for claims in a regulated product, I always:
- Document the dataset version and access date
- Explicitly label results as research benchmarks, not clinical performance
- Re-run key metrics on fresh, local data before filing anything with regulators
Addressing Challenges with Medical Datasets in AI Development
Overcoming Privacy and De-identification Issues in Medical Datasets for AI
Even with public datasets, privacy isn’t “solved.” I keep three rules of thumb:
- Assume re-identification is theoretically possible. Avoid linking public datasets with external sources in ways that might reconstruct identity.
- Respect DUAs to the letter. MIMIC, eICU, and similar resources (MIMIC-III/IV, eICU-CRD) clearly state prohibited uses: I build checks into our data engineering workflows to enforce them.
- Segregate environments. I never mix de-identified public datasets and internal PHI in the same unsecured notebook or S3 bucket.
For HIPAA/GDPR, remember that “de-identified” under US rules doesn’t automatically satisfy GDPR’s definitions of anonymization. EU deployments often require extra legal review even when models were pre-trained on US public data.
Tackling Bias and Ensuring Representativeness in Medical AI Datasets
Bias is where I’ve seen the biggest disconnect between benchmark success and bedside safety.
MIMIC and eICU are US ICU populations: ChestX-ray14 and CheXpert are tertiary-care, mostly academic-hospital chest X-rays. GTEx has its own ancestry imbalances. If I naively train on these and deploy in a different geography or care setting, I’m almost guaranteed to see:
- Shifted calibration (over- or underestimation of risk)
- Performance gaps across age, sex, and race subgroups
- Drift when documentation or imaging protocols differ
My mitigation pattern is:
- Use public datasets for feature/architecture exploration and early benchmarks.
- During internal validation, run stratified performance audits by clinically meaningful subgroups.
- Involve domain experts (clinicians, statisticians, ethics) when interpreting gaps and deciding whether mitigation (re-weighting, re-sampling, additional local data) is adequate.
Finally, if a model can’t clear minimum safety and equity thresholds on local, governed data, I don’t deploy it, regardless of how well it performs on public benchmarks.
Conflict of interest & sponsorship: I have no financial ties to any of the datasets, institutions, or platforms mentioned. All opinions are my own, based on my experience developing and validating medical AI systems as of November 2025.
Disclaimer:
The content on this website is for informational and educational purposes only and is intended to help readers understand AI technologies used in healthcare settings. It does not provide medical advice, diagnosis, treatment, or clinical guidance. Any medical decisions must be made by qualified healthcare professionals. AI models, tools, or workflows described here are assistive technologies, not substitutes for professional medical judgment. Deployment of any AI system in real clinical environments requires institutional approval, regulatory and legal review, data privacy compliance (e.g., HIPAA/GDPR), and oversight by licensed medical personnel. DR7.ai and its authors assume no responsibility for actions taken based on this content.
Frequently Asked Questions About Medical AI Datasets
What are medical AI datasets and why do they matter so much for regulated deployments?
Medical AI datasets are collections of clinical, imaging, or omics data used to train and evaluate models. In regulated environments, they largely determine what you can credibly claim about performance, safety, bias, and robustness to regulators and clinicians—far more than the specific model architecture does.
Which open medical AI datasets are most commonly used for EHR and ICU research?
For EHR-heavy work, MIMIC-III, MIMIC-IV, and the eICU Collaborative Research Database are widely used. They provide de-identified ICU and hospital data, including time-series vitals, labs, medications, and clinical notes, making them ideal for sepsis prediction, mortality risk modeling, note classification, and benchmarking research pipelines.
How should I use medical imaging datasets like ChestX-ray14, CheXpert, and CAMELYON in AI development?
ChestX-ray14 and CheXpert are excellent for pre-training encoders, weakly supervised learning, and benchmarking chest X-ray classifiers. CAMELYON16/17 is foundational for lymph node metastasis detection in pathology. Use them for research, method development, and stress-testing—but always perform final fine-tuning and validation on local, governed clinical data.
Can I use public medical AI datasets like MIMIC or CheXpert in commercial products?
Often yes, but only under strict conditions. You must comply with each dataset’s data use agreement, licensing terms, and privacy constraints, and you still need separate local validation on governed data. Regulators typically expect that clinical claims for commercial products are backed by institution-specific data, not only open benchmarks.
How do I choose the right medical AI dataset for my project and control bias?
Match the dataset to your target setting and modality: MIMIC/eICU for ICU EHR modeling, CheXpert/ChestX‑ray14 for radiology, GTEx/GDC for omics. Recognize their limits—US-centric, ICU-heavy, or narrow disease focus—and run stratified performance audits on your own local data to detect and mitigate subgroup performance gaps.
Past Review:






