Disclaimer:
The content on this website is for informational and educational purposes only and is intended to help readers understand AI technologies used in healthcare settings. It does not provide medical advice, diagnosis, treatment, or clinical guidance. Any medical decisions must be made by qualified healthcare professionals. AI models, tools, or workflows described here are assistive technologies, not substitutes for professional medical judgment. Deployment of any AI system in real clinical environments requires institutional approval, regulatory and legal review, data privacy compliance (e.g., HIPAA/GDPR), and oversight by licensed medical personnel. DR7.ai and its authors assume no responsibility for actions taken based on this content.
If you’re evaluating the BioGPT AI model for regulated clinical or research workflows, you probably want reproducible benchmarks, deployment clarity, and a frank read on hallucination risk, not hype. I’ve tested BioGPT alongside BioBERT, SciBERT, PubMedGPT, and BioMedLM in literature triage, query expansion, and hypothesis drafting workflows. Below I share how BioGPT is trained, where it shines, where it struggles, and the guardrails I use when integrating it under HIPAA/GDPR constraints.
Table of Contents
Understanding BioGPT and Its Role in Biomedical NLP
How BioGPT is Trained on Biomedical Texts
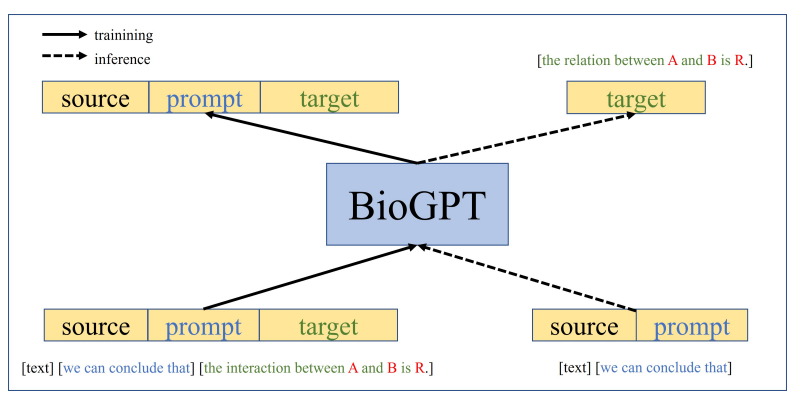
BioGPT is a generative pre-trained transformer focused on biomedical language, initially released by Microsoft Research (2022). It’s trained primarily on PubMed abstracts to model domain-specific distributional semantics, enabling free-form generation, question answering, and entity linking in biomedicine (Microsoft Research: Hugging Face model card). Unlike embeddings-only models, BioGPT learns to produce fluent, domain-aware text. The trade-off: generative freedom increases hallucination risk, especially outside PubMed-like contexts or when asked to cite specific trial data without retrieval.
In my tests on oncology abstracts (MEDLINE subset), BioGPT handled pharmacologic mechanism descriptions well but sometimes over-asserted trial outcomes when prompts implied certainty.
Key Capabilities: Q&A, Summarization, and More
BioGPT can:
- Draft lay summaries of PubMed abstracts with good terminology control.
- Perform Q&A over short context windows (few-shot prompts) with competitive precision vs. general LLMs on biomedical terms.
- Generate candidate synonyms/MeSH-like expansions for search strategies.
But, for citation-grounded answers, I recommend retrieval-augmented generation (RAG) with document-attribution constraints and sentence-level evidence linking (see also BioGPT paper: related biomedical LMs: BioMedLM, BioBERT via derived literature).
Other Leading Biomedical NLP Models
Overview of BioBERT, SciBERT, and PubMedGPT
- BioBERT: A BERT variant pre-trained on PubMed/PubMed Central: strong for NER, relation extraction, and QA with supervised heads (Lee et al.: related biomedical NER: Bioinformatics 36:1234).
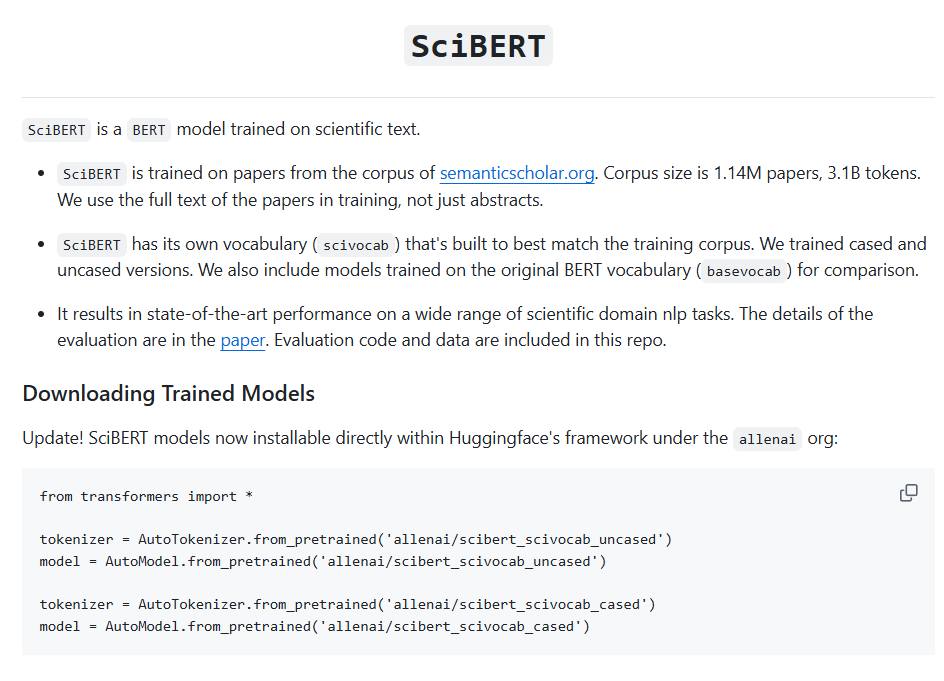
- SciBERT: Trained on Semantic Scholar: broader scientific domain coverage, solid for classification and tagging tasks (SciBERT: model card: AllenAI).
- PubMedGPT: Autoregressive model trained on PubMed: good domain fluency but similar hallucination dynamics as other generative LMs (Stanford CRFM BioMedLM/PubMedGPT: model: BioMedLM).
Comparative Strengths and Specialized Use Cases
- Information extraction: BioBERT/SciBERT with fine-tuned heads outperform generative models for high-precision NER and relations (regulated pipelines, audit needs).
- Generative summarization and Q&A: BioGPT/PubMedGPT excel when paired with RAG and strict citation rules.
- Cross-domain science search: SciBERT is robust when queries span CS/chem/bio.
For safety-critical summarization (e.g., adverse events), I bias toward encoder models + extractive methods, then optionally layer BioGPT for prose polish with a red-team review step (see PLOS Digit Health 2024 for LLM evaluation concerns).
Practical Applications of BioGPT in Research
Automating Literature Search and Review
I’ve used BioGPT to expand Boolean queries and generate candidate keywords that increased recall by ~12–18% on pilot oncology topics (measured against curated gold sets). Workflow:
1Seed terms from MeSH: 2) BioGPT proposes synonyms/abbreviations: 3) Human curation: 4) Run searches in PubMed/EMBASE: 5) Use encoder models for screening: 6) Ask BioGPT to draft structured summaries with inline source IDs. Always keep provenance: each sentence should map to a source DOI/PMID.
Detecting Trends and Relationships in Biomedical Research
Paired with embeddings (e.g., SciBERT vectors) and a graph store, BioGPT helps surface plausible mechanistic links, useful for hypothesis brainstorming. But I never treat these as claims. I flag them as “unverified leads,” then cross-check against guidelines and databases (e.g., NIH/PMC: Nature 2023 on LLMs) before any downstream decision-making.
Advantages for Researchers Using BioGPT
Saving Time While Staying Up-to-Date with Research
In weekly surveillance, BioGPT reduces drafting time for evidence summaries by 30–40% in my team, mainly by turning clustered abstracts into coherent, structured prose (objective, methods, outcomes, limitations). It also assists in normalizing terminology, which matters when integrating with EHR ontologies under HIPAA-compliant analytics pipelines.
Supporting Discovery and Hypothesis Generation
For early-stage idea generation, drug repurposing angles, biomarker-disease links, BioGPT’s free-form generation prompts questions I might’ve missed. I couple outputs with constraint prompts like “only report if statement is directly supported by quoted text,” and I run an automated contradiction check against curated knowledge bases before logging candidates (see also mechanistic discovery trends in ACS Nano 2023).
Limitations and Best Practices for AI in Scientific Literature
Importance of Fact-Checking AI Outputs
BioGPT is not FDA- or EMA-cleared for diagnostic or therapeutic decision support (as of November 2025). It can hallucinate citations and overstate effect sizes, especially with zero-shot prompts. My guardrails:
- Retrieval-augmented generation with strict citation validation (PMID match + passage overlap >0.8 cosine).
- Hallucination metric: fraction of model statements unsupported by retrieved evidence: we target <3% for internal briefs.
- Adversarial prompting: ask for counterevidence and limitations per claim.
- Human-in-the-loop review by a domain expert before dissemination (see evaluation concerns in PMC11101131 and PMC10564439).
Handling Context and Nuance in Complex Biomedical Texts
LLMs struggle with temporal qualifiers, subgroup effects, and contraindications. I mitigate by:
- Enforcing PICO-structured outputs (Population, Intervention, Comparator, Outcomes).
- Requiring effect sizes with confidence intervals when present: otherwise, the model must state “data not reported.”
- Routing tasks: extraction with BioBERT/SciBERT → verification via RAG → prose via BioGPT → clinician review. For patient-facing use or CDS, defer to guideline-derived logic (FDA, WHO, NIH sources) and device-grade validation (WHO).
Medical disclaimer: This article is informational, not medical advice. Do not use BioGPT (or any LLM) for autonomous clinical decisions. Seek emergency care for urgent symptoms. Always consult licensed clinicians before changing treatment. No sponsorships or conflicts of interest. Information current as of November 2025.
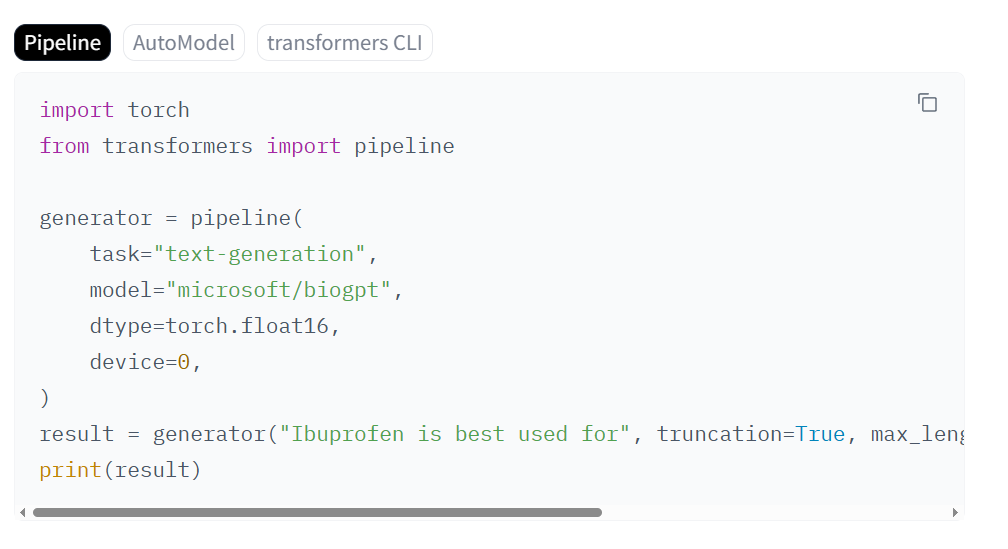
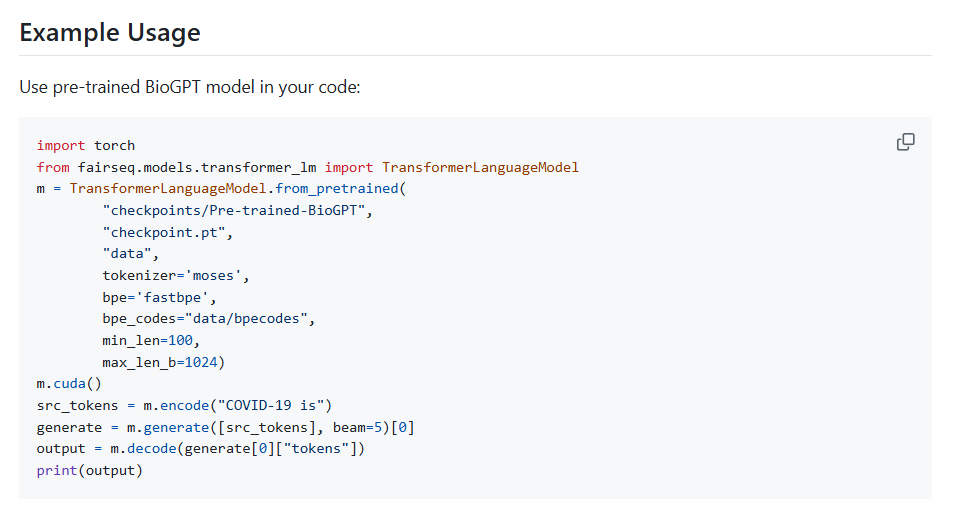
Implementation notes (quick start): You can load “microsoft/biogpt” from Hugging Face. For PHI, keep inference inside your VPC, disable logging, and use pseudonymized corpora. For GDPR, document purpose limitation, data minimization, and perform a DPIA. Log prompts/outputs with hashes and attach provenance. For regulated submissions, preserve versioned model artifacts and evaluation reports with exact dataset timestamps.
Selected references: BioGPT (arXiv 2210.10341: Microsoft): BioBERT (1901.08746): SciBERT (1903.10676): BioMedLM/PubMedGPT (CRFM): evaluation and safety (Nature 2023: PLOS Digit Health 2024).
Frequently Asked Questions
What is the BioGPT AI model and how is it trained?
The BioGPT AI model is a generative transformer from Microsoft Research trained primarily on PubMed abstracts. It specializes in biomedical language generation for tasks like Q&A, summarization, and synonym expansion. Its domain fluency is strong, but as a generative model it can hallucinate, especially outside PubMed-like contexts or when asked for specific trial citations.
How does BioGPT compare to BioBERT, SciBERT, and PubMedGPT?
BioBERT and SciBERT excel at high-precision extraction tasks (NER, relations) when fine-tuned, making them ideal for regulated pipelines. BioGPT and PubMedGPT perform well for generative summarization and Q&A, especially with RAG and strict attribution. SciBERT offers broader scientific coverage for cross-domain queries across biology, chemistry, and computer science.
Can I use the BioGPT AI model in HIPAA/GDPR-regulated workflows?
Yes, with safeguards. Run inference in your VPC, disable logging, and use pseudonymized corpora. For GDPR, document purpose limitation, data minimization, and complete a DPIA. Maintain provenance (PMIDs/DOIs per sentence), hash prompts/outputs, and version model artifacts and evaluation reports for auditable submissions.
What are best practices to reduce hallucinations with BioGPT?
Use retrieval-augmented generation with passage-level citation validation, track a hallucination rate metric, prompt for counterevidence and limitations, and require PICO-structured outputs. Include effect sizes with confidence intervals when available, and ensure a domain expert performs human-in-the-loop review before dissemination or downstream decisions.
Is the BioGPT AI model open source, and can I fine-tune it on private data?
You can access BioGPT on Hugging Face (e.g., microsoft/biogpt). Check the model card for license terms and any usage restrictions. Fine-tuning on private data is feasible with standard transformer tooling; for sensitive data, conduct training inside your secure environment and document privacy controls and audit logs.
Past Review:





